Usage
inc_overlap(
.data,
.fun,
.step = 1000,
.n.steps = 10,
.downsample = FALSE,
.bootstrap = NA,
.verbose.inc = TRUE,
...
)Arguments
- .data
The data to be processed. Can be data.frame, data.table::data.table, or a list of these objects.
Every object must have columns in the immunarch compatible format. immunarch_data_format
Competent users may provide advanced data representations: DBI database connections, or a list of these objects. They are supported with the same limitations as basic objects.
Note: each connection must represent a separate repertoire.
- .fun
Function to compute overlaps. e.g.,
morisita_index.- .step
Either an integer or a numeric vector.
In the first case, the integer defines the step of incremental overlap.
In the second case, the vector encodes all repertoire sampling depths.
- .n.steps
Integer. Number of steps if
.stepis a single integer. Skipped if ".step" is a numeric vector.- .downsample
If TRUE then performs downsampling to N clonotypes at each step instead of choosing the top N clonotypes.
- .bootstrap
Set NA to turn off any bootstrapping, set a number to perform bootstrapping with this number of tries.
- .verbose.inc
Logical. If TRUE then shows the output from the computation process.
- ...
Other arguments passed to
.fun.
Examples
# \dontrun{
data(immdata)
ov <- repOverlap(immdata$data, "inc+overlap", .step = 100, .verbose.inc = FALSE, .verbose = FALSE)
vis(ov)
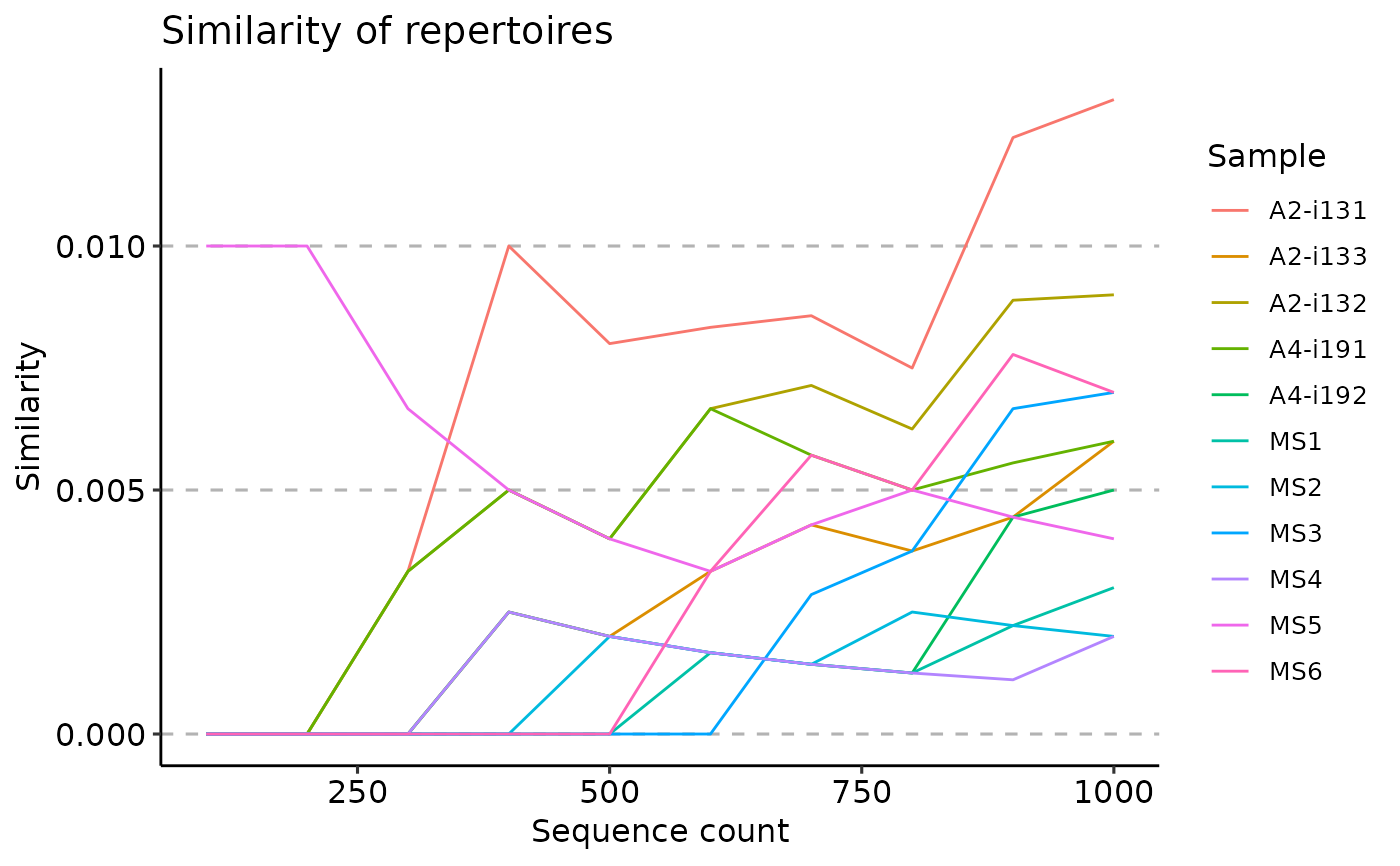 # }
# }