An utility function to analyse the immune receptor gene usage
(IGHD, IGHJ, IDHV, IGIJ, IGKJ, IGKV, IGLJ, IGLV, TRAJ, TRAV, TRBD, etc.)
and statistics. For gene details run gene_stats().
Arguments
- .data
The data to be processed. Can be data.frame, data.table::data.table, or a list of these objects.
Every object must have columns in the immunarch compatible format. immunarch_data_format
Competent users may provide advanced data representations: DBI database connections,or a list of these objects. They are supported with the same limitations as basic objects.
Note: each connection must represent a separate repertoire.
- .gene
A character vector of length one with the name of the gene you want to analyse of the specific species. If you provide a vector of different length, only the first element will be used. The string should also contain the species of interest, for example, valid ".gene" arguments are "hs.trbv", "HomoSapiens.TRBJ" or "macmul.IGHV". For details run
gene_stats().- .quant
Selects the column with data to evaluate. Pass NA if you want to compute gene statistics at the clonotype level without re-weighting. Pass "count" to use the "Clones" column to weight genes by abundance of their corresponding clonotypes.
- .ambig
An option to handle ambiguous gene assigments, e.g., "TRAV1,TRAV2".
Pass "inc" to include all possible gene segments, so "TRAV1,TRAV2" is counted as a different gene segment.
Pass "exc" to exclude all ambiguous gene assignments, so "TRAV1,TRAV2" is excluded from the resultant gene table.
We recommend to turn it on by passing "inc" (turned on by default). You can exclude data for the cases where there is no clear match for gene, include it for every supplied gene, or pick only first from the set. Set it to "exc", "inc" or "maj", respectively.
- .type
Set the type of data to evaluate: "segment", "allele", or "family".
- .norm
If TRUE then return proportions of genes. If FALSE then return counts of genes.
Value
A data frame with rows corresponding to gene segments and columns corresponding to the input samples.
Examples
data(immdata)
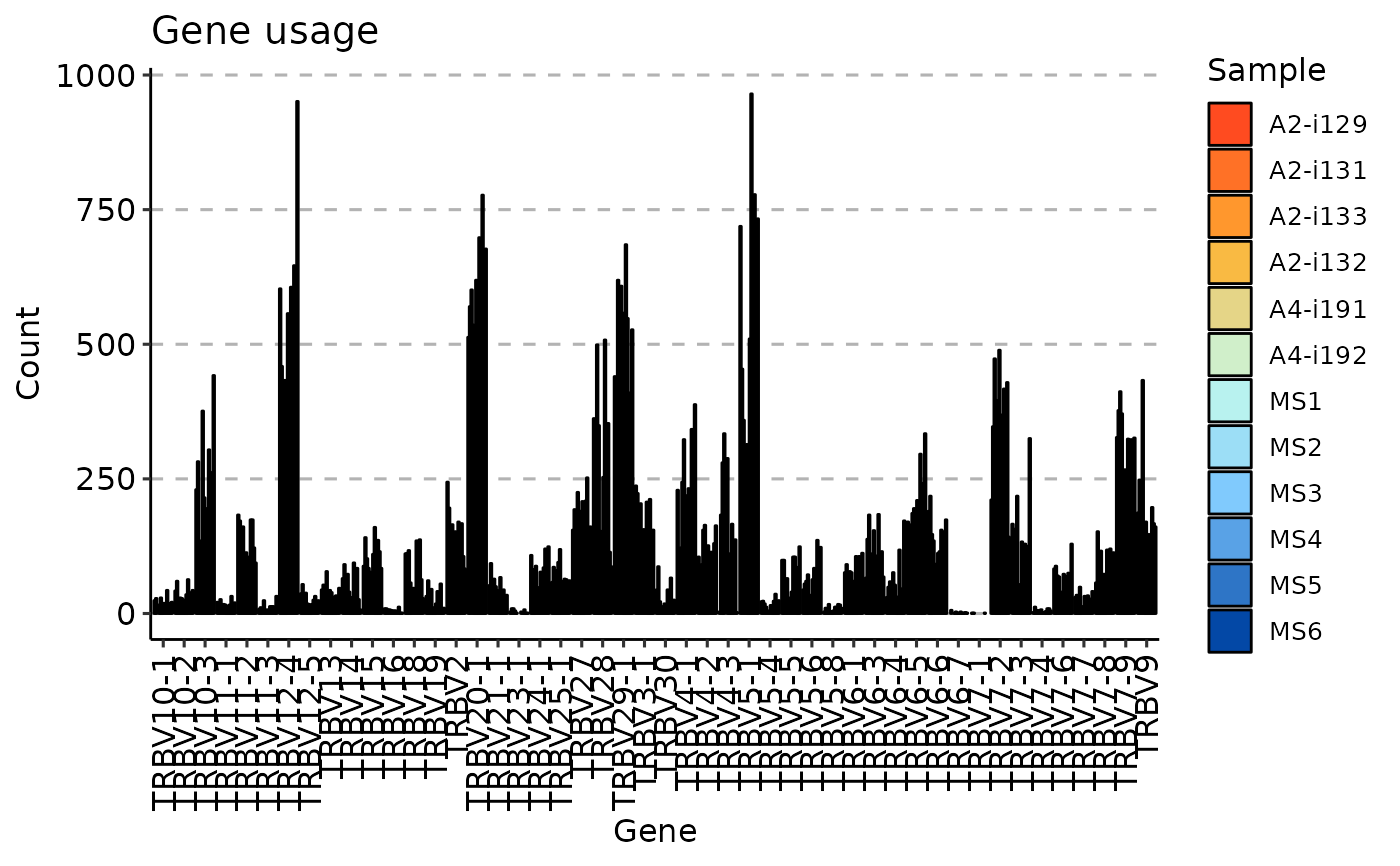
gu <- geneUsage(immdata$data)
vis(gu)
#> Using Names as id variables
#> Warning: The `size` argument of `element_line()` is deprecated as of ggplot2 3.4.0.
#> ℹ Please use the `linewidth` argument instead.
#> ℹ The deprecated feature was likely used in the immunarch package.
#> Please report the issue at <https://github.com/immunomind/immunarch/issues>.
#> Warning: The `size` argument of `element_rect()` is deprecated as of ggplot2 3.4.0.
#> ℹ Please use the `linewidth` argument instead.
#> ℹ The deprecated feature was likely used in the immunarch package.
#> Please report the issue at <https://github.com/immunomind/immunarch/issues>.
#> Warning: Removed 15 rows containing missing values or values outside the scale range
#> (`geom_bar()`).